How to Conduct Localized Proteomics of Microscopic Regions

Dr. Eleanor Drummond
Read BioEleanor completed her PhD in Neuroscience at the University of Western Australia in Perth, Australia. She studied the role of beta amyloid and sex hormones in Alzheimer’s disease. She was then recruited to New York University, School of Medicine as a postdoc. She is currently a Research Assistant Professor of Neurology at NYU where she uses proteomics to understand the pathogenesis of Alzheimer’s disease.
ClosePrecise and lightning-fast spectral Fluorescence Lifetime Imaging at video rate integrated in a high-end confocal microscope
In this tutorial, you will learn:
- How to perform isolated proteomics of microscopic regions of interest
- How to use this technique with FFPE tissue
- How proteomics can help you understand molecular mechanisms in disease
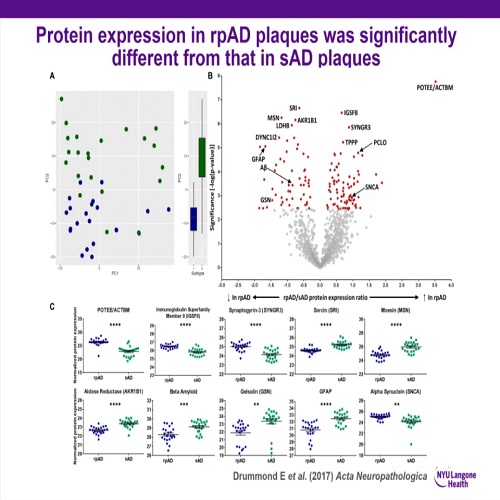
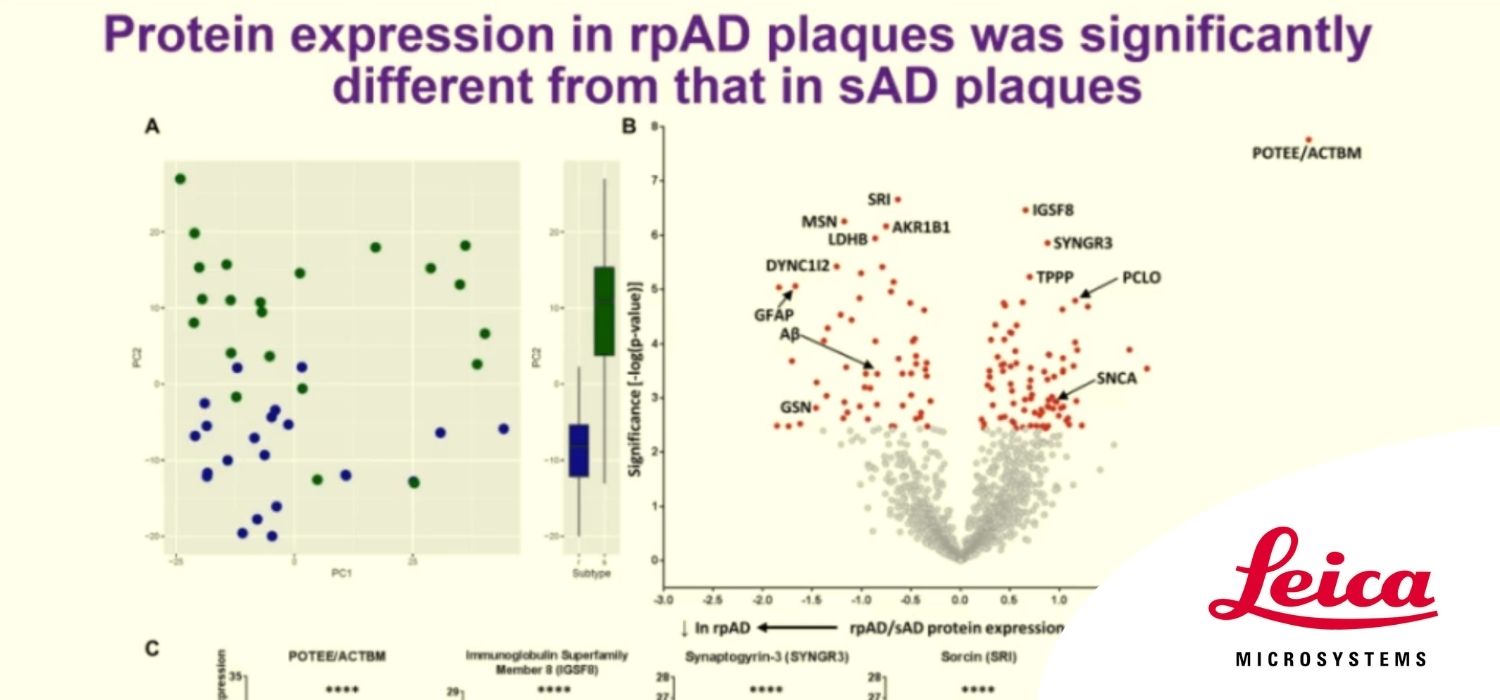
We recently developed a technique that allows localized proteomics of microscopic regions of interest, such as specific cell types or neuropathological features of disease. This technique uses laser capture microdissection (LCM) to isolate regions/cells of interest followed by label-free quantitative mass spectrometry (LC-MS). Importantly, we optimized this technique to use formalin-fixed paraffin embedded (FFPE) tissue, so that archived human tissue specimens collected at autopsy could be used. This is a particular advantage of our methodology, as the vast majority of human tissue specimens are FFPE blocks, which are currently an underutilized, but exceptionally valuable resource for medical research. We have successfully used this technique to analyze the proteome of neuropathological features that define Alzheimer’s disease (amyloid plaques and neurofibrillary tangles), as well as specific populations of neurons that are vulnerable in AD. Going forward, the use of localized proteomics has the potential to greatly increase our understanding of the molecular mechanisms that underlie AD. More broadly, this technique could be used to analyze regions or cells of interest isolated from any FFPE tissue, and therefore could be widely used to examine disease pathogenesis across a broad spectrum of diseases.